To Issue 129
Citation: Lee E, “Evolving Analytical Methods: Advances in Molecule Characterisation”. ONdrugDelivery Issue 129 (Feb 2022), pp 60–63.
Eliza Lee discusses some of the modern analytical techniques available for the characterisation of modern biologics, and how CDMOs equipped with these techniques are able to support pharmaceutical development.
“Executing these finicky assessments accurately can prove challenging for even the most experienced teams. Fortunately, advances in the design and development of commercially available analytical products and kits have enabled highly accurate analyses with rapid run times.”
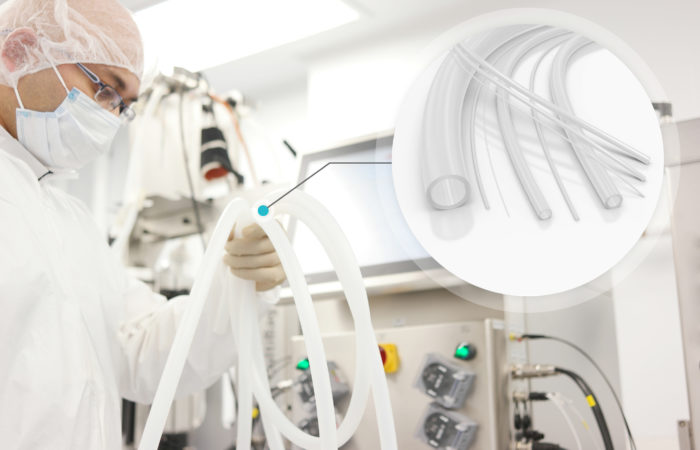
Comprehensive analytical assessments of structure and function are essential in order to compliantly bring a molecule to market. With recent technological advances, it is possible to characterise molecules and their interactions more thoroughly than ever before. Moreover, these robust analytical methods are often required by regulatory bodies as part of a successful filing for commercial manufacturing. It is therefore essential for CDMOs to remain at the forefront of analytical technologies (Figure 1).

Figure 1: It is key for CDMOs to be able to carry out the robust analyses required by regulators to help pharma partners successfully see their products to market.
Researchers and developers are under constant pressure to deliver results from increasingly thorough analytical assessments and analyses during drug development. Characterising the structure and activity of today’s more complicated biotherapeutic molecules requires highly sensitive informatics systems and analytical technologies. To meet this demand for insightful and high-fidelity data, researchers and developers must use numerous advanced analytical technologies to deliver sophisticated assessments in the most effective and cost-efficient way.
UNDERSTANDING THE MOLECULE – BEGIN WITH ACTIVITY AND FUNCTION
Assessment of molecular function is necessary for determining whether the desired cellular or immune responses are likely to be stimulated in vivo. Therefore, functional assays play an essential role in understanding a molecule’s potency.
The assays used to analyse molecular function must reflect cellular and immune system responses, which may involve complex, sophisticated feedback-controlled interactions and reactions. As such, executing these finicky assessments accurately can prove challenging for even the most experienced teams. Fortunately, advances in the design and development of commercially available analytical products and kits have enabled highly accurate analyses with rapid run times.
ELISA – How Potent is the Molecule?
Best practice for researchers during early-phase development is to use binding assays, such as enzyme-linked immunosorbent assays (ELISAs), also known as enzyme immunoassays (EIAs), to examine the potency of their biological molecule. This is frequently the preferred initial assay when starting biomolecular analysis.1
The ELISA is a plate-based technique designed to detect and quantify soluble substances that are used to elucidate interactions between highly specific antibodies and antigens indirectly. The assay facilitates conditions where the interaction of interest immobilises the participating molecules onto the wells of a microplate. After washing and removing unbound components, a substrate is added to the microplate wells, which reacts with reporter enzymes linked to antibodies. This reaction produces a quantitative signal which is proportional to the dose-dependent interaction.
The use of an ELISA to determine potency has many benefits, including high specificity, sensitivity and efficiency, as well as a simple procedure. Another advantage is the shorter timelines required for development and qualification, which can help to accelerate investigational new drug (IND) filing times. However, depending on the mode of action (MoA) of the molecule, some projects may necessitate additional cell-based assays.
“The actives and compounds in a biologic formulation are likely to have contact with a broad range of materials during manufacture.”
Cell-based Assays – What Does the Molecule do to Cell Systems?
Antigen-specific ELISAs allow for analysis of interactions under highly controlled conditions but may not accurately represent interactions in vivo. Molecules that have an MoA that can affect the functioning of entire systems within a cell may require cell-based assays to suitably assess their impact.
There are many cell-based assays that can be deployed. Assessment of cell viability (by determining the ratio of live and dead cells) and cell proliferation (by examining how the molecule impacts cell division over time) are two common examples. Other key cell-based assays for analysing molecule activity include cell signalling, cytotoxicity and cell apoptosis.
Cell-based assays can also be used to measure the effect of anticancer drugs. Depending on the type of analysis used, the results of this assay could act to support technical transfer as well as support chemistry, manufacturing and controls (CMC) development.2
UNDERSTANDING THE MOLECULE – DETERMINE THE SAFETY PROFILE
Analytical tools are essential for ascertaining whether a molecule, or other component present within the finished biopharmaceutical, is safe for use. Biological and chemical contaminants must be removed, or confirmed to be present only at a level that will not affect the patient. Without these assessments, untested drugs could cause toxicity, carcinogenic reactions, immunogenic reactions or other adverse events.
PCR: Have the Biological Contaminants Been Removed?
Polymerase chain reaction (PCR) assays are an effective and efficient method of making many copies, or “amplifying”, small segments of DNA.3 As high concentrations of DNA are not abundant in biological systems, using DNA as a substrate in analytical assays has previously been nearly impossible. With the advent of PCR in 1993, low concentrations of DNA can be amplified to levels suitable for use in in vitro assay analyses (Figure 2). Laboratory and clinical techniques, including diagnosis of genetic disorders, fingerprinting DNA and detection of bacteria or viruses, are now all supported by PCR.

Figure 2: PCR techniques enable DNA concentrations to be amplified to suitable levels for analysis.
Quantitative PCR can be used to amplify specific segments of contaminant DNA that may be present at specific stages of the molecule manufacturing process, such as residual host cell DNA (HCD). The reaction is run over a specific timeframe, after which the amount of amplified DNA can be determined, which is proportional to the amount of contaminant present. Both the WHO and US FDA have stringent guidelines for the concentration of acceptable HCD.
Reverse-transcriptase PCR (RT-PCR) is commonly used to identify the presence of viruses. Some viruses carry their genetic information in the form of RNA rather than DNA, meaning conventional quantitative PCR cannot be used to detect their presence. Instead, RT-PCR, which includes a step where RNA is converted to DNA by the enzyme reverse transcriptase, is necessary. Viral clearance analysis is required to ensure that process purification steps have removed potentially contaminating viruses that could be introduced during manufacturing or from raw materials.
Analysing the Presence of Leachables and Extractables:
Have Chemical Contaminants Been Removed?
Most biopharmaceuticals are in liquid form throughout the manufacturing process, and remain that way until they are aseptically filled and finished into primary containers for parenteral delivery. As a result, the actives and compounds in a biologic formulation are likely to have contact with a broad range of materials during manufacture.
All materials and surfaces that have been in contact with the formulation must be analysed to determine if any interactions have occurred. Some extractable and leachable compounds can impact the quality of therapeutic biological proteins by altering their physicochemical properties. Their presence in the formulation could also have the potential to cause toxic, carcinogenic or immunogenic effects in patients.
Researchers must seek a full and comprehensive understanding of the presence and possible effects of all extractables and leachables in the final formulation. Without achieving this early in development, stability and contamination risks could severely disrupt the drug development and manufacturing programme.
“The importance of operational optimisation in analytical method
development is increasing in parallel with the advancement of analytical technologies.”
Capillary Electrophoresis: What is the Size and Charge of the Molecule?
In the development and manufacturing of many biological molecules, it is important to determine the molecule’s stability and identify physiochemical modifications. Previously, techniques such as sodium dodecyl sulphate–polyacrylamide gel electrophoresis (SDS-PAGE) and isoelectric focusing (IEF) were used to assess size variation (associated with stability) and charge variation (associated with physiochemical modifications) respectively. These methods are time consuming and labour intensive.
Over the past decade, more accurate and cost-effective capillary electrophoresis (CE) methods have replaced SDS-PAGE and IEF. These include capillary electrophoresis SDS (CE-SDS) and capillary IEF (cIEF), both of which are capable of producing more robust and reproducible results compared with their predecessors.
NOVEL ANALYTICS: HOW CAN WE ANALYSE NOVEL MOLECULES?
Poorly characterised complex molecules, such as fusion proteins, recombinant proteins and multi-specific antibodies, pose novel analytical challenges. Evaluating critical quality attributes (CQAs) is particularly difficult for complex molecules. Their analysis therefore necessitates the use of novel techniques.
Better characterisation of novel molecules can be achieved by optimising commonly applied analytics such as CE methods and high-performance liquid chromatography (HPLC). Careful optimisation of run conditions can ensure reproducibility and robustness in methods determining molecule characteristics.
Optimisation is also required in methods for quantifying surfactants, such as polysorbate 80 (PS80). Although these techniques have improved in recent times, driven in part by the interest regulators have taken in providing more formulation characterisation information in their guidance, optimisation is needed to raise their robustness and reproducibility.
Glycan analysis is another important technique that is now frequently required for providing information on more complex proteins, such as in vivo half-life, effector function and immunogenicity. Glycosylation can be challenging to study because it occurs heterogeneously; branching and single sites with multiple glycan structures present are commonly observed. Although quantitative glycosylation can provide reproducible and robust data, it is labour intensive and time-consuming. On the other hand, high-throughput glycan analysis is much faster, but may only provide qualitative results. It is best practice to choose which glycolysis analysis to use based on data needs. In particular, considerations should be given to how the target molecule’s protein activity will be affected by glycosylation at each stage of the process.
THE IMPACT OF OPTIMISING ANALYSIS
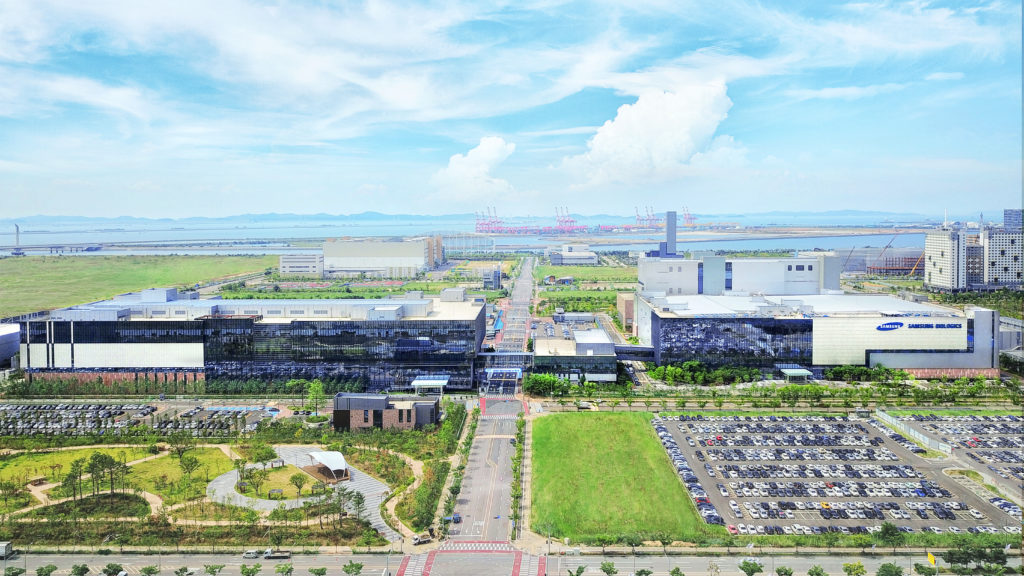

Figure 3: The Samsung Biologics site in Incheon, Republic of Korea, includes all the facilities necessary to support pharma using robust platform methods and analytical technology.
CDMOs, equipped with robust platform methods, are now able to assess product quality more effectively than ever before (Figure 3). The importance of operational optimisation in analytical method development is increasing in parallel with the advancement of analytical technologies. Moreover, this approach facilitates timely, cost-effective discovery, particularly when high-throughput analysis is integrated into analytical platform methods. As a result, biopharmaceutical drugs can move quickly through clinical trials to patients.
REFERENCES
- “Overview of ELISA”. ThermoFisher Scientific website, accessed Dec 2020.
- Goldberg R, “Analytical Methods for Cell Therapies: Method Development and Validation Challenges”. BioProcess International, Jun 2021.
- “Polymerase Chain Reaction (PCR) Fact Sheet”. National Human Genome Research Institute, Aug 2020.